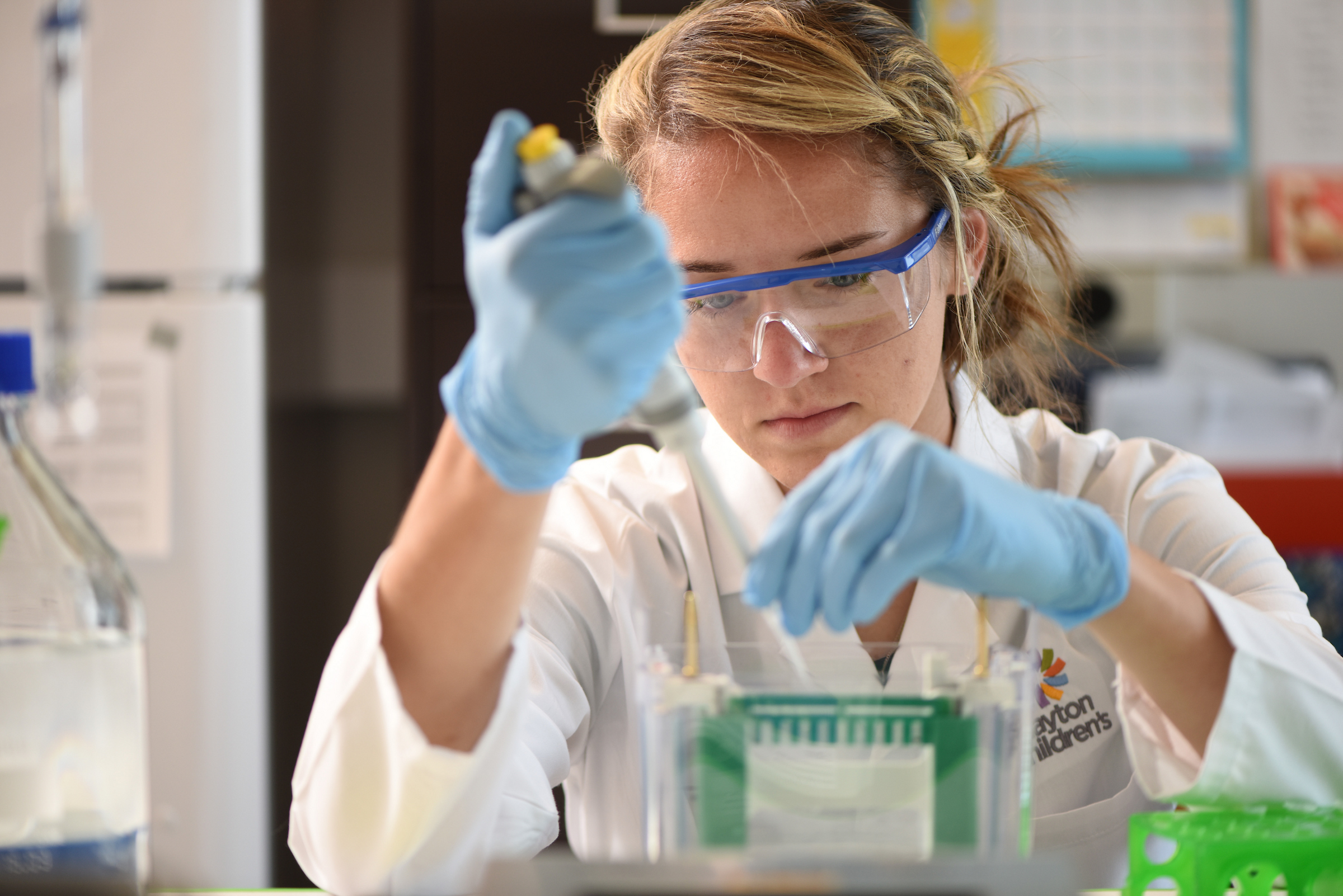
The Living Biobank at Dayton Children’s
The Living Biobank at Dayton Children’s advances pediatric brain tumor research through imaging studies, molecular data and global collaboration. Get involved today.


Dayton Children’s researchers are working to better understand how brain tumors differ from one another, and even within a single tumor. By studying the molecular features of tumor cells grown in different microenvironments, our team explores how those differences influence treatment response.
These findings may explain why small biopsy samples sometimes fail to capture the full picture of a tumor’s biology. They also help us identify new imaging and metabolic markers that could allow clinicians to predict diagnosis and prognosis non-invasively.
bridging imaging and biology
Our research connects in vivo tumor imaging—how a tumor behaves inside the body—with in vitro cell culture studies that analyze growth and metabolism in controlled environments.
This combined approach helps reveal how the tumor’s microenvironment affects its sensitivity to therapy.
Using artificial intelligence and deep learning, we conduct advanced image analysis that integrates clinical, biologic, and genetic data. Together, these tools allow us to create more accurate preclinical tumor models and explore imaging techniques that may guide therapy decisions without surgical intervention.
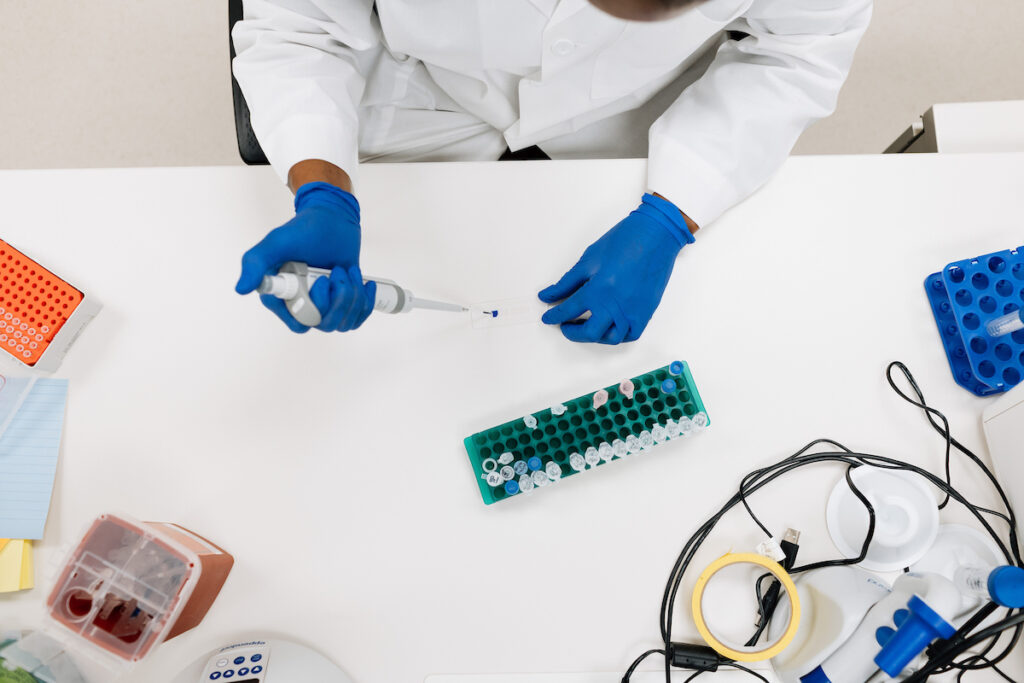


research collaboration and impact
With a mission to improve the lives of children with brain tumors, the Living Biobank brings together a multi-institutional network of pediatric neurosurgeons, oncologists, radiologists, pathologists, physicists, mathematicians, and biochemists.
By combining data, expertise, and technology, we are building a translational research platform that accelerates the development of safer and more precise treatments for children.
background and significance
Pediatric brain tumors have surpassed leukemia as the leading cause of cancer-related death in children. Treatment resistance often results from intra-tumoral heterogeneity—variations within a single tumor that remain poorly understood.
Traditional models, limited biological samples, and reliance on adult data have slowed pediatric progress. Current personalized approaches require high risk, invasive biopsies and rarely accessible genomic testing. Non-invasive techniques for diagnosis, prognosis, and design of tailored therapies are not available.
Our long-term goal is to develop non-invasive imaging methods that can identify tumor characteristics and guide therapeutic decisions. By linking automated image analysis to molecular and metabolic data from experimental models, the Living Biobank aims to provide clinicians with practical tools to support individualized care and next-generation preclinical research.
specific aims
The brain tumor microenvironment is heterogeneous, with variable features of hypoxia, acidosis, nutrient depletion, and stromal and immune cell signaling. These have clear effects on the hallmarks of tumor growth, invasion, and migration, but have not been well characterized in the context of in vivo imaging and preclinical cell culture models.
The specific aims of our research will test the overarching hypothesis that microenvironmental stressors along with stromal and immune components can.
We are collaborating with Dr. Kristen Yeom at Stanford University on application of advanced imaging techniques to the study of pediatric brain tumors. Our work has described how patterns of microscopic water diffusion and blood perfusion can be used to distinguish histopathologic features that are otherwise ambiguous on conventional imaging. We have defined prognostically distinct tumor subtypes that correlate with clinical behavior. We also described a phenomenon of low blood perfusion in a specific type of high-grade glioma that is suggestive of a hypoxic state, and has formed the basis of our current in vitro experiments evaluating the effects of hypoxia on epigenetic modifications in patient-derived tumor cultures.
We have created a shared imaging server and data use agreement between Dayton Children’s Hospital and Stanford University, allowing us to share brain tumor data for validation of a novel deep learning model of approximately 250 separate MRI features, based on computer vision tasks that are completely independent of human input.
We have a similar collaboration with Dr. Jason Parker at Indiana University, also in the field of advanced imaging. Our work has also focused on machine learning for integrating and classifying MRI features of brain tumors, based on the rationale that tumor heterogeneity can contribute to treatment failure. This is a separate, second strategy of combining multiple imaging modalities into a single predictive framework, known as “nosologic imaging analysis,” and includes correlation of data collected from intraoperative stereotactic navigation images to biological specimens obtained during tumor resection.
With financial support from the Gala of Hope, we established a Tissue Bank and culture facility at Dayton Children’s Hospital, now with a current inventory of over 20 cultured central nervous system tumors linked to highly detailed clinical and imaging data elements. We have recently become satellite members of the Children’s Brain Tumor Tissue Consortium. Work is in progress to expand our culture program to include orthotopic xenograft mouse models and lentoviral infection for immortalization of low-grade tumors prone to in vitro senescence.
In our laboratory at the Wright State University Neuroscience Engineering Collaboration Building, we are studying the effects of microenvironmental stress on autopsy-derived diffuse intrinsic pontine glioma (DIPG) cells obtained from Stanford University and VU Medical Center in Amsterdam.
research highlights
Recent studies from the Living Biobank and collaborating institutions explore the biology and imaging of pediatric brain tumors to inform future treatments.
Portions of DIPG tumors have low blood flow and poor oxygenation, which activates proteins known as hypoxia-inducible factors (HIF). This potentially causes them to behave more aggressively.
In these studies we found that individual DIPG tumors had distinct responses to low oxygen. In general HIF proteins appeared to be activated regardless of how much oxygen was present, impacting tumor growth and metabolism.
Waker, C.A., Keoni, C., Schurko, B., Brown, T.L., Lober, R.M. Hypoxia-inducible factors regulate diffuse intrinsic pontine glioma growth in normoxic culture. Presented at the International Society for Pediatric Neuro-Oncology, Denver, CO, July 1, 2018. https://doi.org/10.1093/neuonc/noy059.160
Waker, C.A., Shahin, M., Kamian, A., Lober, Z.C., Lober, R.M. Differential hypoxic response in human DIPG cell lines. Presented at the Society for Neuro-Oncology, New York, NY, June 15–16, 2017. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5475014/
This is pioneering work spearheaded by our collaborator Dr. Jason Parker at Indiana University. It is a mathematical approach to classifying types of brain tumor tissue based on clinical imaging. Machine learning was applied to understand how each piece of information from an MRI scan can contribute to a computer’s ability to classify the tumor tissue into specific categories, which may be used in the future to help with diagnosis, prognosis, and treatment planning.
Parker, J.G., Diller, E.E., Lober, R.M. Quantifying individual and collective prediction accuracy of MR contrasts for glioma tissue compartment classification. Presented at the International Society for Magnetic Resonance in Medicine / European Society for Magnetic Resonance in Medicine and Biology, Paris, France, June 19, 2018.
samples – The Living Biobank
Stay up-to-date with the latest insights from Dayton Children’s Hospital. We’re always working to share helpful, real-world content for families navigating pediatric allergy and immunology care.
tumor specimens
- 55MNU1482: cerebellar pilocytic astrocytoma
- 59FNG4928: Recurrent parietal low grade glioneural tumor with FGFR1 duplication
- 63MNU3810: Supratentorial ependymoma
- 64MNU3453: Subependymal giant cell astrocytoma
- 65MNJ3624: Recurrent/residual DNET
- 66FNU1623: Cerebellar pilocytic astrocytoma
- 67MNK6150: Juvenile angiofibroma
- 68MNM3154: Metastatic rhabdomyosarcoma to the brain
- 70FNU5453: Intraventricular meningioma
- 71MNU5238: Right occipital isomorphic astrocytoma
- 72MNU698: Cerebellar pilocytic astrocytoma with gangliocytic differentiation
- 73MNU4107: Posterior fossa pilocytic astrocytoma
- 75MNU2341: Posterior fossa epidermoid cyst
- 79MNO1378: Anaplastic medulloblastoma
- 80FND5577: Recurrent disseminated pilomyxoid astrocytoma
- 81FNQ3716: Recurrent primitive neuroectodermal tumor
- 82MNU3290: Dorsal medullary pilocytic astrocytoma
- 87MNV781: Intraventricular high grade glioma
- 88MHW5715: Glioblastoma
- 88MNV5550: High grade glioma
- 89FNU995: Posterior fossa pilocytic astrocytoma
- 90FNL18538: Meningioma
- 91FNL26461: Meningioma
- 92FNU5047: Pleomorphic xanthoastrocytoma
- 93FNU1676: Ependymoma
- 95FNV2166: High grade glioma
- 96FNUX9006: Recurrent sphenoid wing meningioma
- 98MNU5971: Pilocytic astrocytoma
- 100MNU4934: Glioblastoma
- 103FNU4702: Meningioma
- 105FNU1372: Epidermoid cyst
- 107M..1897: Embryonal rhabdomyosarcoma
- 109FU912: Astroblastoma
- 110M_3620: Histiocytosis
- 113MMO5961: Medulloblastoma
- 114MU370: ATRT
- 115FU3837: Pilocytic astrocytoma
- 21FNB4389: Recurrent schwannoma: Hurler syndrome
- 74MNU726: Post-radiation ganglioneuroma
- 77MNN2895: Plexiform neurofibroma of the face
- 78MNA6318: Spinal intramedullary neurofibroma
- 84FNR7626: Myxopapillary ependymoma of the filum terminale
- 85MNA6520: Spinal schwannoma
- 94MNA6445: Thoracic spinal schwannoma
- 97MNU4174: Intramedullary spinal cord GBM
- 56MNF4628: Diffuse cranial osteoma
- 23MCH2940: Chiari I malformation
- 60FCH4028: Chiari I malformation
- 61MCH5781: Chiari I malformation
- 62FCH4005: Chiari I malformation
- 76FCH1894: Chiari I malformation
- 104FCH2598: Chiari I malformation
- 106MCH461: Chiari I malformation
- 112FH6841: Chiari I malformation
- 116MH3583: Chiari I malformation
tissue from craniofacial specimens
- 22MCA81: Sagittal synostosis
- 22MCA116: Sagittal synostosis
- 34MCA115: Sagittal synostosis
- 57MCB111: Metopic synostosis
- 58FCC150: Coronal synostosis
- 69MCD164: Bicoronal synostosis; Muenke syndrome
- 83MCA124: Sagittal and metopic synostosis
- 99MC_48: Sagittal synostosis
- 101FCA153: Sagittal synostosis
- 102MCA…: Sagittal synostosis
- 86FCE519: Orofacial cleft and hypertelorism
- 108ML7005: Temporal lobectomy
- 111ML_2707: Temporal lobectomy
our partners at the Living Biobank
Our research is bridging the gap between pediatric neurosurgeons, pediatric oncologists, physicists, mathematicians, radiologists, pathologists, biochemists and many others within Dayton Children’s and the Neuroscience Institute. We combine artificial intelligence, clinical, biologic, and genetic data and an unprecedented level of advanced image analysis in efforts to understand and develop treatments for tumors.
Since its founding with support from the Gala of Hope Foundation, the Living Biobank has grown into a global network of research and clinical partners. Support from local foundations and national collaborations enables our team to collect, study and share biospecimens and data that advance pediatric brain tumor research worldwide.
The work continues through the love, generosity and support of our partners, including many families who have been affected by brain tumors. Together we are making progress that will serve generations to come.
- Gala of Hope Foundation – established the Living Biobank in 2016 with a $198,870 gift
- John French Estate
- Hartzell Norris Charitable Trust
- Neils and Ruth Lundgard Foundation
- Ohio Space Grant Consortium
- Boonshoft School of Medicine Medical Student Research Grant
- Wright State University Honors Program Research Scholarship
- Mayfield Education and Research Foundation (MERF) Spark Grant
- DIPG/DMG Collaborative – $69,600 grant (2019)
- Barry & Denise Johnson Foundation – $133,500 raised through the annual Ezra Hartke Race for Hope
- Speedway Children’s Charities – funded purchase of the Invenio NIO Laser Imaging System
- Children’s Brain Tumor Network (Children’s Hospital of Philadelphia)
- International Pediatric Brain Tumor Atlas
- Childhood Cancer Model Atlas (Hudson Institute of Medical Research, Australia)
- Washington University in St. Louis
- University of Calgary
- Hackensack Meridian Health – Center for Discovery & Innovation
- Seattle Children’s Hospital
- Stanford University
- Indiana University
- Kettering Health (Dr. Neil Patel and neurosurgery team)
- Gift From a Child national program
support the search for a cure
Your gift will change the life of a child with brain cancer. The Living Biobank at Dayton Children’s is on a mission to eliminate pediatric brain cancer. Support from our caring community is essential to supplying the Living Biobank with the very best tools and resources, so they can carry on with their important work.
get involved with the Living Biobank
Collaboration is at the heart of the Living Biobank. We partner with biologists, physicians, physicists, and mathematicians, research professionals and more. In addition to our current areas of need, there are many volunteer opportunities for individuals of all ages and experiences.

Clinical projects at Dayton Children’s are available to medical students interested in contributing to pediatric brain tumor research.
Qualified undergraduate students can participate in independent study projects leading to laboratory experiments or honors projects at Wright State University.
Middle and high school students can design school science fair projects or mentor younger students to raise awareness about pediatric brain tumors. High school students may also organize volunteer activities, work on in silico experiments, mobile applications or web-based projects.
Even our youngest supporters can help make a difference by creating artwork that promotes brain tumor research or cards and gifts for our patients. These small acts of creativity inspire hope and help share the mission of the Living Biobank with others.
Your gift will change the life of a child with brain cancer. The Living Biobank at Dayton Children’s is on a mission to eliminate pediatric brain cancer. Our world-class research team is determined to improve the lives of kids right here in the Miami Valley and around the world. Support from our caring community is essential to supplying the Living Biobank with the very best tools and resources, so they can carry on with their important work. Your gift will help provide life-changing and life-saving cancer care for kids.
areas of current need
- Clinical data entry
- CRISPR/Cas9 gene editing
- Python programming language for machine learning
- Image analysis & automated segmentation
- Bioinformatics
- Scientific & medical illustration
- Augmented reality presentations
- Community organizations & fundraising
- School outreach, teaching & student mentorship
- Making the world a better place
make a difference
Help advance pediatric brain tumor research by contributing your skills, creativity or research proposal to the Living Biobank team.
research & innovate with us
Together, we can transform the future of pediatric care. Whether through participating in research, collaborating on innovative programs, or supporting our mission, there are many ways to get involved.
